The cores listed below are located at Harvard Medical School, affiliated institutions, or other Harvard Schools.
Bioinformatics/Data Analytics
Big Data Analytics Core
The Big Data Analytics Core services include predictive analytics with biomedical data including cohort studies and EHR data, text mining and automated knowledge extraction, real-world evidence, as well as design and analysis of modern clinical trials.
BioGrids
BioGrids provides innovative computational resources, scientific software, and educational programs. We curate a library of 500+ biosciences applications to support researchers across disciplines with collections for high-throughput sequencing, proteomics, genomics, visualization, and related tools, and provide access to these tools through a preconfigured software environment.
Core Director - Jason Key
Operations Director - Michelle Ottaviano
Core Website - https://www.biogrids.org/
BPF NGS Genomics Core Facility
The BPF NGS Genomics Core Facility provides state of the art resources and services including NextGen Sequencing on the Illumina platform, NGS Sample Preparation (for a variety of DNA and RNA applications), Single Cell Analysis on the 10X Genomics Chromium platform, DNA and RNA Quality Assessment, Sanger DNA Sequencing, DNA/RNA isolation and purification, Oligonucleotide Ordering, qPCR Assays, and Reagents & Supplies Ordering (via a staffed stock room in the NRB and an automated 24/7 stock room on the HMS Quad).
Core Director - Robert Steen
Core Website - https://genome.med.harvard.edu/
For additional information on the Biopolymer Facility, visit their page on the Catalyst Directory
Center for Computational Biomedicine
CCB provides cutting edge computational capabilities, data analysis, and data integration technologies to support research and education across HMS. Our multi-disciplinary team of computational and quantitative scientists develops shared data and analytic resources to serve a broad constituency and provides expertise and management for project collaborations with HMS investigators. CCB also hosts a variety of events (i.e., workshops, guided learnings, town halls, seminars) to enhance computational education for HMS students, trainees, research staff, and faculty.
Drosophila RNAi Screening Center
The mission of the DRSC is to develop and optimize new molecular genetic technologies for use in vivo in Drosophila (TRiP technologies), as well as for use in Drosophila, mosquito, and other insect cell lines (DRSC technologies). In addition, we provide access to information, protocols, data and a suite of online tools via our website, and develop research resources such as new fly stocks and modified insect cell lines. We also have state-of-the-art facilities for in vivo Drosophila genetics and insect tissue culture, and provide support for small- and large-scale insect cell screening projects using RNAi or CRISPR-Cas technologies.
Core Director - Stephanie Mohr
Core Website - http://fgr.hms.harvard.edu/
HMS imaging cores website: https://microscopy.hms.harvard.edu
Harvard Chan Bioinformatics Core
In an effort to provide bioinformatics analysis services and training to the HMS Community, the HMS Tools and Technology Program has provided additional support and resources to the HSPH Bioinformatics Core (HBC). HBC provides expertise in areas such as array analysis, next-gen sequencing (NGS) and functional analysis. Their NGS support includes epigenetics, transcriptomics and re-sequencing studies. HBC works together with research computing groups on all aspects of data management.
Core Director - Shannan Ho-Sui
Core Website - http://bioinformatics.hms.harvard.edu/
Image Management Core at HMS
The Harvard Medical School Image Management Core is a new service of HMS Research IT Solutions to help researchers manage image data and metadata. The IMC was created in response to the growing complexity and difficulty of managing research image data. The first major offering of the IMC will be a shared installation of the OMERO image management platform. The HMS IT department will install and maintain an OMERO server for use by on-quad HMS researchers. This effort builds upon the successful use of the OMERO platform by the HMS LINCS project. Initial funding for this comes from the Tools and Technology Committee, HMS IT, and the Harvard Program in Therapeutic Science (HiTS).
Core Director- Jay Copeland
Core Website - http://imc.hms.harvard.edu/
Nascent Transcriptomics Core
The mission of the Nascent Transcriptomics Core (NTC) is to offer the community a resource for the analysis of nascent transcription, providing new insights into gene regulation and enabling highly-sensitive identification of regulatory regions such as enhancers.
We offer consultation on experimental design as well as library construction services for Start-seq, PRO-seq, and TT-seq. In addition, the NTC looks forward to expanding these offerings as we develop and optimize new nascent RNA sequencing methods in partnership with the HMS community.
Core Contact - Seth Goldman
Core Website - https://ntc.hms.harvard.edu/
Research Computing
The Research Computing (RC) Core is a set of billable services provided by Research Computing and HMS IT. The goal of the RC Core is to promote deeper collaboration across the greater Harvard biomedical research ecosystem. This includes improved capabilities and performance by establishing more transparent and sustainable IT services for our research community.
Research Computing will be providing the following billable services via the RC Core:
-
High Performance Computing:
- The O2 Cluster, which accommodates diverse requirements and workflows for HMS-affiliated researchers.
-
Storage Options:
- Active – intended for storing research data that is frequently accessed, modified, or computed against which includes Compute and Collaborations storage solutions.
- Standby - is leveraged for infrequently accessed data that is still directly available for reference, retrieval, or analysis.
Core Director - Neil Coplan
Core Website - https://it.hms.harvard.edu/rc/core
Email - rccore@hms.harvard.edu
SBGrid Advanced Research Computing (ARC) and Software Consortium
SBGrid Consortium provides innovative computational resources, scientific software, and educational programs to researchers in the global structural biology community. We curate a library of 500+ scientific applications to support all phases of macromolecular structure determination with software collections for electron microscopy, X-ray crystallography, NMR, computational chemistry, and structure prediction, and provide access to these tools through a preconfigured software environment.
SBGrid Advanced Research Computing (ARC) provides research computing support to structural biology laboratories in the Boston area, assisting with the design and maintenance of research computing infrastructure, including workstation and server hardware purchase recommendations.
Core Director - Jason Key
Operations Director - Michelle Ottaviano
ARC Website - https://sbgrid.org/corewiki/Home
Consortium Website - https://www.sbgrid.org/
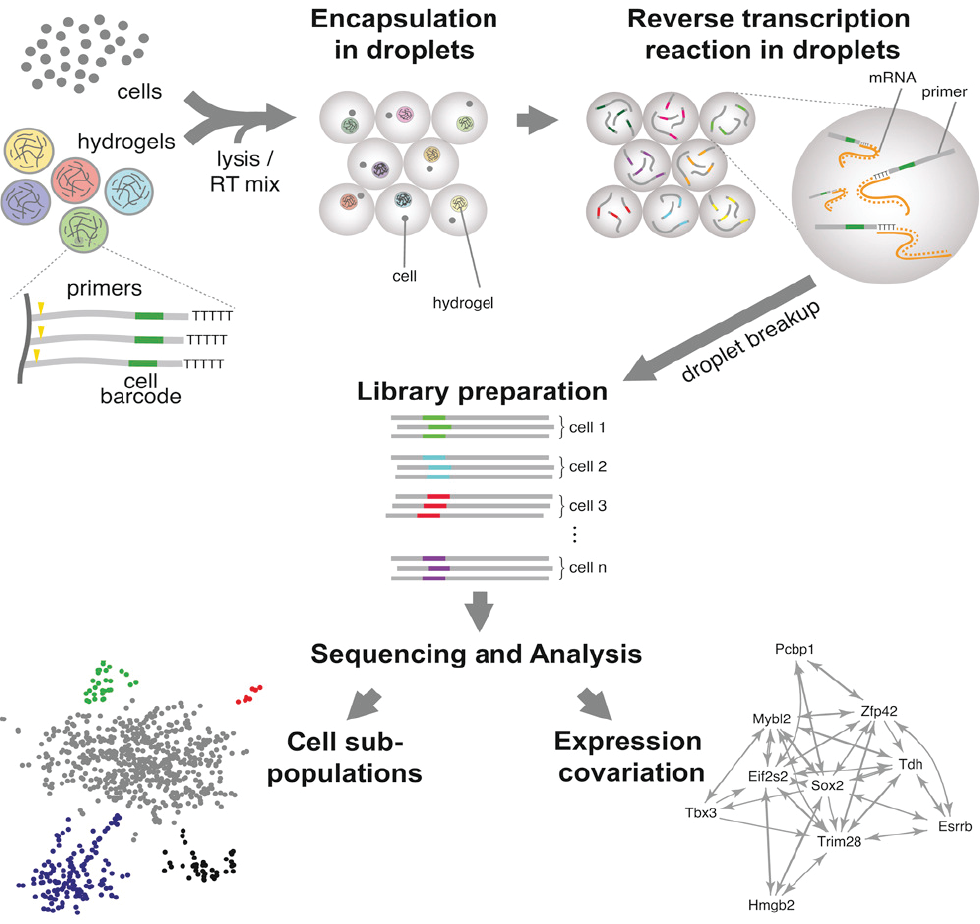
Single Cell Core at HMS
This Core supports single cell sequencing and provides resources for sequencing the transcriptomes individual cells and provides scientific advising for optimal experimental design. The Core is partnering with the Harvard Chan Bioinformatics Core to offer support from experimental design through to data analysis.
Core Director - Mandovi Chatterjee
Core Website - https://singlecellcore.hms.harvard.edu/